

|
|||
|
|||
A plant
PTB-like protein is translocated in the phloem |
|||
Jeri Brandom 1 , Byung-Chun Yoo 2 , Nien-Chen Huang 1 and William J. Lucas 1 . |
|||
| 1 Section of Plant Biology, Division of Biological Sciences, University of California, Davis, California. 2 Current address: Delaware Biotechnology Institute, Newark, Delaware. | |||
There is increasing evidence that plants use non-cell autonomous movement of ribonucleoproteins (RNPs) to coordinate regulation of developmental and physiological processes (Haywood et al. , 2002). Systemic RNA signaling is consistent with the long-distance transmission of RNA interference and the ability of translocated aberrant transcripts to alter developmental phenotypes. In animals, RNA localization is a key determinant of developmental fate. RNA localization also occurs during embryogenesis and polarized cell growth in plants (Okita and Choi, 2002). Many RNA-binding proteins (RBPs) involved in cytoplasmic localization of RNA in animals have been identified and several of these, by sequence similarity, have apparent homologs in plants. However, little is known about proteins that may mediate long-distance RNA signaling in plants. Polypyrimidine tract-binding protein (PTB, also known as hnRNP I) is an RBP involved intracellular RNA localization in animals (Cote et al., 1999). It is a multi-functional RBP with four loosely conserved RNA-recognition motifs (RRM). PTB binds to C/U rich polypyrimidine tracts and can be purified from nuclear extracts with poly(U) affinity chromatography. In addition to RNA-localization, PTB plays an important role in the intracellular processing of RNA, including nuclear export of mRNA, alternative splicing of pre-mRNA, mRNA stabilization, regulation of cap-independent translation, and poly(A) site cleavage. PTB proteins have been identified in many animal systems and may be ubiquitous in metazoans. Sequence similarity indicates the presence of PTB homologs in several plant species. The Arabidopsis genome contains both PTB homologs and homologs of proteins that antagonize the activity of PTB in 3′-splice site selection. However, no plant PTB has yet been characterized. In order to identify components that mediate systemic signaling, we have conducted a screen of pumpkin phloem exudate for the presence of poly(U) RNA-binding proteins. We identified a 50-kD phloem localized protein, Cucurbita maxima RNA-binding protein, 50kD ( CmRBP50 ) with sequence similarity to animal PTBs. Considering the ability of PTB to mediate RNA localization and influence the ultimate pattern of gene expression, it is of great interest that there is a PTB-like protein present in the phloem. Heterograft analysis indicates that this protein is translocated long-distance in the phloem. Possible conserved PTB-like activity of CmRBP50, as well as a potential role for CmRBP50 in selective systemic trafficking of transcripts or long-distance signaling, is being investigated.
|
|||
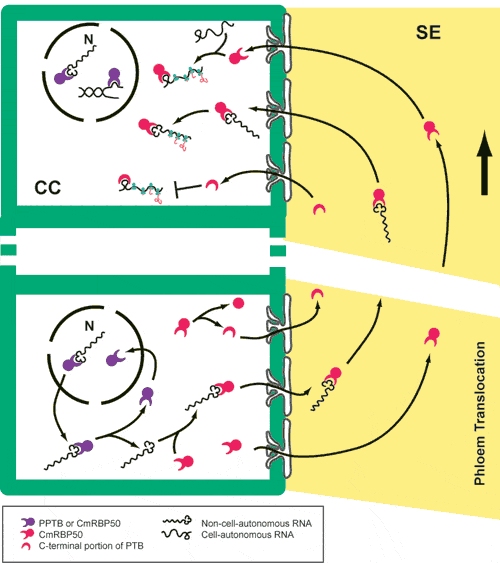 |
What are the possible functions of CmRBP50? If CmRBP50 has conserved PTB-like activity, it may play a role in Alternative splicing of pre-mRNA, poly (A) site cleavage, nuclear export of mRNA, cytoplasmic localization of mRNA, cap-independent translation, or regulation of transcription. As an RNA-binding protein that is translocated long distance in the plant, it may function in long-distance localization of mRNA. |
||
This work is supported by a grant from the DOE Division of Energy Biosciences (DE-FG03-94ER20134). |
|||
|
|||
References Haywood, V., Kragler, F., Lucas, W.J. (2002) Plasmodesmata: pathways for protein and ribonucleoprotein signaling. Plant Cell supplement 2002: S303-S325. Okita, T.W., and Choi, S.-B. (2002) mRNA localization in plants: targeting to the cell's cortical region and beyond. Curr. Opin. Plant Biol. 5: 553-559. Cote, C. A., Gautreau, D., Denegre, J.M., Kress, T.L., Terry, N.A, Mowry, K.L. (1999) A Xenopus protein related to hnRNP I has a role in cytoplasmic RNA localization. Mol. Cell. 4: 431-437. |
|||
